Immunometabolism
The immunometabolism Core is a hub within the institute for analysis of metabolism using the instrumentation and methodology that has been established at LJI as well as through partnerships across UC facilities. All of the instruments and approaches are available as a service to LJI scientists, but also (as capacity permits) to outside users.
Thomas Riffelmacher, Ph.D.
Services
Seahorse metabolic flux assays
The Extracellular Flux Analyzer (XF-96 Seahorse Biosciences/Agilent Technologies) is available at LJI and allows for measurements of oxygen consumption rates (OCR) and extracellular acidification rates (ECAR) in a 96-well plate format in intact cells, permeabilized cells, spheroids, and isolated mitochondria. XF Analyzers perform compound addition and mixing, label-free analytical detection, and automatic measurement of OCR and ECAR in real time. The most common application is live cell respirometry, providing live information on metabolic flux and cellular energetic phenotype, ATP rates, and mitochondrial activity in isolated (immune) cells. Isolated mitochondria from animal tissues or cell lines can also be used for mechanistic studies.
Single cell resolution metabolism
We also perform nutrient/metabolite consumption assays using labelled glucose, amino acid or fatty acid molecules in vivo (injection + tracing on single cell level across organs) or in vitro. These assays provide further information on the metabolic programs and nutrient preference in immune cells at single cell resolution, and can be complemented with single-cell resolution assays on mitochondrial mass and -potential, lipid stores and other metabolic organelles as well as immune markers to create UMAP maps of metabolic states of single cells.
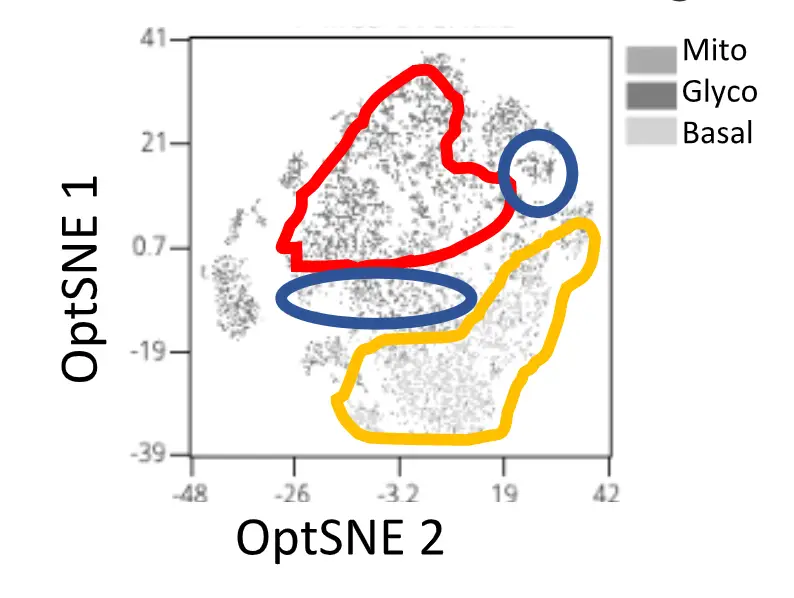
Sub-cellular metabolism + metabolomic analysis
We provide sample preparation, acquisition and analysis by transmission electron microscopy (TEM) to study metabolic organelle morphology in partnership with the UCSD Facility (Mitochondrial ultrastructure, lipid stores, autophagosomes, …). 105 cells minimum/sample. This can reveal mitochondrial shape changes, including cristae structure, and their association with lipid droplets and other relevant organelles.
For whole metabolome study, 13C Carbon labelling and tracing can be performed in isolated immune cells (105 cells minimum/sample). Analysis by mass spectrometry reveals isotope incorporation across glycolytic metabolites, pentose phosphate pathway metabolites, citric acid cycle, ribose and nucleotide metabolites, and fatty acids. These data reveal particular metabolic phenotypes at the level of individual metabolites and facilitate molecular mechanistic understanding but also inform global pathway preference and activity.
Seahorse metabolic flux assays
The Extracellular Flux Analyzer (XF-96 Seahorse Biosciences/Agilent Technologies) is available at LJI and allows for measurements of oxygen consumption rates (OCR) and extracellular acidification rates (ECAR) in a 96-well plate format in intact cells, permeabilized cells, spheroids, and isolated mitochondria. XF Analyzers perform compound addition and mixing, label-free analytical detection, and automatic measurement of OCR and ECAR in real time. The most common application is live cell respirometry, providing live information on metabolic flux and cellular energetic
phenotype, ATP rates, and mitochondrial activity in isolated (immune) cells. Isolated mitochondria from animal tissues or cell lines can also be used for mechanistic studies.
Contact
For all scheduling requests, please contact Tom Riffelmacher.

